Table of contents
- 1. Introduction to Biology2h 42m
- 2. Chemistry3h 40m
- 3. Water1h 26m
- 4. Biomolecules2h 23m
- 5. Cell Components2h 26m
- 6. The Membrane2h 31m
- 7. Energy and Metabolism2h 0m
- 8. Respiration2h 40m
- 9. Photosynthesis2h 49m
- 10. Cell Signaling59m
- 11. Cell Division2h 47m
- 12. Meiosis2h 0m
- 13. Mendelian Genetics4h 44m
- Introduction to Mendel's Experiments7m
- Genotype vs. Phenotype17m
- Punnett Squares13m
- Mendel's Experiments26m
- Mendel's Laws18m
- Monohybrid Crosses19m
- Test Crosses14m
- Dihybrid Crosses20m
- Punnett Square Probability26m
- Incomplete Dominance vs. Codominance20m
- Epistasis7m
- Non-Mendelian Genetics12m
- Pedigrees6m
- Autosomal Inheritance21m
- Sex-Linked Inheritance43m
- X-Inactivation9m
- 14. DNA Synthesis2h 27m
- 15. Gene Expression3h 20m
- 16. Regulation of Expression3h 31m
- Introduction to Regulation of Gene Expression13m
- Prokaryotic Gene Regulation via Operons27m
- The Lac Operon21m
- Glucose's Impact on Lac Operon25m
- The Trp Operon20m
- Review of the Lac Operon & Trp Operon11m
- Introduction to Eukaryotic Gene Regulation9m
- Eukaryotic Chromatin Modifications16m
- Eukaryotic Transcriptional Control22m
- Eukaryotic Post-Transcriptional Regulation28m
- Eukaryotic Post-Translational Regulation13m
- 17. Viruses37m
- 18. Biotechnology2h 58m
- 19. Genomics17m
- 20. Development1h 5m
- 21. Evolution3h 1m
- 22. Evolution of Populations3h 52m
- 23. Speciation1h 37m
- 24. History of Life on Earth2h 6m
- 25. Phylogeny2h 31m
- 26. Prokaryotes4h 59m
- 27. Protists1h 12m
- 28. Plants1h 22m
- 29. Fungi36m
- 30. Overview of Animals34m
- 31. Invertebrates1h 2m
- 32. Vertebrates50m
- 33. Plant Anatomy1h 3m
- 34. Vascular Plant Transport1h 2m
- 35. Soil37m
- 36. Plant Reproduction47m
- 37. Plant Sensation and Response1h 9m
- 38. Animal Form and Function1h 19m
- 39. Digestive System1h 10m
- 40. Circulatory System1h 57m
- 41. Immune System1h 12m
- 42. Osmoregulation and Excretion50m
- 43. Endocrine System1h 4m
- 44. Animal Reproduction1h 2m
- 45. Nervous System1h 55m
- 46. Sensory Systems46m
- 47. Muscle Systems23m
- 48. Ecology3h 11m
- Introduction to Ecology20m
- Biogeography14m
- Earth's Climate Patterns50m
- Introduction to Terrestrial Biomes10m
- Terrestrial Biomes: Near Equator13m
- Terrestrial Biomes: Temperate Regions10m
- Terrestrial Biomes: Northern Regions15m
- Introduction to Aquatic Biomes27m
- Freshwater Aquatic Biomes14m
- Marine Aquatic Biomes13m
- 49. Animal Behavior28m
- 50. Population Ecology3h 41m
- Introduction to Population Ecology28m
- Population Sampling Methods23m
- Life History12m
- Population Demography17m
- Factors Limiting Population Growth14m
- Introduction to Population Growth Models22m
- Linear Population Growth6m
- Exponential Population Growth29m
- Logistic Population Growth32m
- r/K Selection10m
- The Human Population22m
- 51. Community Ecology2h 46m
- Introduction to Community Ecology2m
- Introduction to Community Interactions9m
- Community Interactions: Competition (-/-)38m
- Community Interactions: Exploitation (+/-)23m
- Community Interactions: Mutualism (+/+) & Commensalism (+/0)9m
- Community Structure35m
- Community Dynamics26m
- Geographic Impact on Communities21m
- 52. Ecosystems2h 36m
- 53. Conservation Biology24m
15. Gene Expression
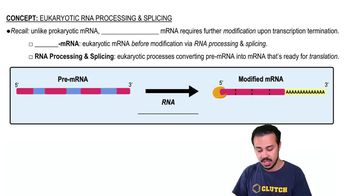
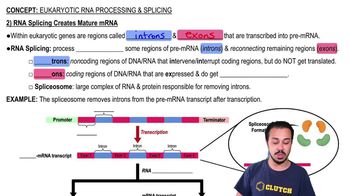
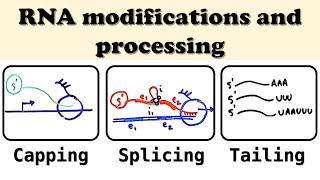
Eukaryotic RNA Processing and Splicing
Problem 4`
Textbook Question
Compared with mRNAs that have a cap and tail, predict what will be observed if a eukaryotic mRNA lacked a cap and poly(A) tail.
a. The primary transcript would not be processed properly.
b. Translation would occur inefficiently.
c. Enzymes on the ribosome would add a cap and poly(A) tail.
d. tRNAs would become more resistant to degradation.
 Verified step by step guidance
Verified step by step guidance1
Understand the role of the cap and poly(A) tail in eukaryotic mRNA. The 5' cap is added to the beginning of the mRNA during processing and helps protect the mRNA from degradation, assists in ribosome binding during translation, and facilitates nuclear export. The poly(A) tail, added to the 3' end, also protects the mRNA from degradation and aids in translation efficiency.
Analyze the options provided in the question. Option (a) refers to improper processing of the primary transcript, which is not directly related to the absence of a cap and tail since these modifications occur after transcription. Option (b) suggests inefficient translation, which is plausible because the cap and tail are critical for ribosome binding and stability. Option (c) implies that enzymes on the ribosome would add the cap and tail, which is incorrect because these modifications occur in the nucleus, not on the ribosome. Option (d) mentions tRNAs, which are unrelated to mRNA processing and translation.
Focus on the biological consequences of lacking a cap and poly(A) tail. Without these modifications, the mRNA is more susceptible to degradation by exonucleases, and the ribosome may struggle to efficiently bind and initiate translation.
Consider the process of translation initiation in eukaryotes. The 5' cap is recognized by initiation factors that recruit the ribosome, and the poly(A) tail interacts with proteins that stabilize the mRNA and enhance translation. Without these features, translation efficiency would be significantly reduced.
Conclude that the most likely observation is inefficient translation (option b), as the absence of the cap and poly(A) tail compromises mRNA stability and ribosome binding, which are essential for effective protein synthesis.
 Verified video answer for a similar problem:
Verified video answer for a similar problem:This video solution was recommended by our tutors as helpful for the problem above
Video duration:
48sPlay a video:
Was this helpful?
Key Concepts
Here are the essential concepts you must grasp in order to answer the question correctly.
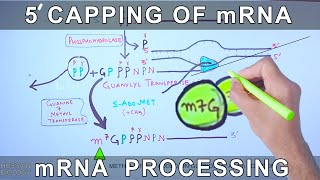
mRNA Capping and Polyadenylation
In eukaryotic cells, mRNA undergoes capping and polyadenylation, which are essential modifications. The 5' cap protects the mRNA from degradation and assists in ribosome binding during translation. The poly(A) tail at the 3' end also stabilizes the mRNA and facilitates its export from the nucleus. Without these modifications, mRNA is more susceptible to degradation and less efficient in translation.
Recommended video:
Guided course

2) mRNA Protection in the Cytoplasm
Translation Efficiency
Translation efficiency refers to how effectively ribosomes synthesize proteins from mRNA. A properly capped and polyadenylated mRNA enhances ribosome recognition and binding, leading to more efficient translation. If an mRNA lacks these features, it may not be recognized by the ribosome, resulting in slower or incomplete translation processes.
Recommended video:
Guided course

Making Sense of Ecosystem Production & Efficiency
mRNA Degradation Mechanisms
Eukaryotic mRNAs are subject to degradation by cellular enzymes, particularly when they lack protective features like the 5' cap and poly(A) tail. These modifications help prevent exonucleases from degrading the mRNA. Without them, the mRNA is more vulnerable to degradation, which can lead to reduced protein synthesis and cellular function.
Recommended video:
Guided course

2) mRNA Protection in the Cytoplasm

 2:32m
2:32mWatch next
Master Eukaryotic RNA Processing and Splicing with a bite sized video explanation from Jason
Start learningRelated Videos
Related Practice