Table of contents
- 1. Introduction to Biology2h 42m
- 2. Chemistry3h 40m
- 3. Water1h 26m
- 4. Biomolecules2h 23m
- 5. Cell Components2h 26m
- 6. The Membrane2h 31m
- 7. Energy and Metabolism2h 0m
- 8. Respiration2h 40m
- 9. Photosynthesis2h 49m
- 10. Cell Signaling59m
- 11. Cell Division2h 47m
- 12. Meiosis2h 0m
- 13. Mendelian Genetics4h 44m
- Introduction to Mendel's Experiments7m
- Genotype vs. Phenotype17m
- Punnett Squares13m
- Mendel's Experiments26m
- Mendel's Laws18m
- Monohybrid Crosses19m
- Test Crosses14m
- Dihybrid Crosses20m
- Punnett Square Probability26m
- Incomplete Dominance vs. Codominance20m
- Epistasis7m
- Non-Mendelian Genetics12m
- Pedigrees6m
- Autosomal Inheritance21m
- Sex-Linked Inheritance43m
- X-Inactivation9m
- 14. DNA Synthesis2h 27m
- 15. Gene Expression3h 20m
- 16. Regulation of Expression3h 31m
- Introduction to Regulation of Gene Expression13m
- Prokaryotic Gene Regulation via Operons27m
- The Lac Operon21m
- Glucose's Impact on Lac Operon25m
- The Trp Operon20m
- Review of the Lac Operon & Trp Operon11m
- Introduction to Eukaryotic Gene Regulation9m
- Eukaryotic Chromatin Modifications16m
- Eukaryotic Transcriptional Control22m
- Eukaryotic Post-Transcriptional Regulation28m
- Eukaryotic Post-Translational Regulation13m
- 17. Viruses37m
- 18. Biotechnology2h 58m
- 19. Genomics17m
- 20. Development1h 5m
- 21. Evolution3h 1m
- 22. Evolution of Populations3h 52m
- 23. Speciation1h 37m
- 24. History of Life on Earth2h 6m
- 25. Phylogeny2h 31m
- 26. Prokaryotes4h 59m
- 27. Protists1h 12m
- 28. Plants1h 22m
- 29. Fungi36m
- 30. Overview of Animals34m
- 31. Invertebrates1h 2m
- 32. Vertebrates50m
- 33. Plant Anatomy1h 3m
- 34. Vascular Plant Transport1h 2m
- 35. Soil37m
- 36. Plant Reproduction47m
- 37. Plant Sensation and Response1h 9m
- 38. Animal Form and Function1h 19m
- 39. Digestive System1h 10m
- 40. Circulatory System1h 57m
- 41. Immune System1h 12m
- 42. Osmoregulation and Excretion50m
- 43. Endocrine System1h 4m
- 44. Animal Reproduction1h 2m
- 45. Nervous System1h 55m
- 46. Sensory Systems46m
- 47. Muscle Systems23m
- 48. Ecology3h 11m
- Introduction to Ecology20m
- Biogeography14m
- Earth's Climate Patterns50m
- Introduction to Terrestrial Biomes10m
- Terrestrial Biomes: Near Equator13m
- Terrestrial Biomes: Temperate Regions10m
- Terrestrial Biomes: Northern Regions15m
- Introduction to Aquatic Biomes27m
- Freshwater Aquatic Biomes14m
- Marine Aquatic Biomes13m
- 49. Animal Behavior28m
- 50. Population Ecology3h 41m
- Introduction to Population Ecology28m
- Population Sampling Methods23m
- Life History12m
- Population Demography17m
- Factors Limiting Population Growth14m
- Introduction to Population Growth Models22m
- Linear Population Growth6m
- Exponential Population Growth29m
- Logistic Population Growth32m
- r/K Selection10m
- The Human Population22m
- 51. Community Ecology2h 46m
- Introduction to Community Ecology2m
- Introduction to Community Interactions9m
- Community Interactions: Competition (-/-)38m
- Community Interactions: Exploitation (+/-)23m
- Community Interactions: Mutualism (+/+) & Commensalism (+/0)9m
- Community Structure35m
- Community Dynamics26m
- Geographic Impact on Communities21m
- 52. Ecosystems2h 36m
- 53. Conservation Biology24m
19. Genomics
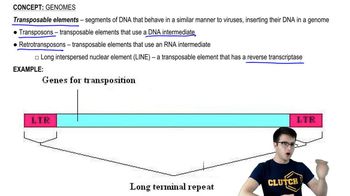
Genomes
Problem 6`
Textbook Question
Consider the validity of the following statements about genome editing. Select True or False for each statement.
T/F Cas proteins work as endonucleases.
T/F sgRNA is used by bacterial cells to detect which DNA to cut.
T/F Homologous recombination is always used to join pieces of broken DNA.
T/F It is possible to modify genes as well as disrupt them by genome editing.
 Verified step by step guidance
Verified step by step guidance1
Understand the role of Cas proteins: Cas proteins are part of the CRISPR system and function as endonucleases, which means they can cut DNA at specific sites. This is a key aspect of genome editing.
Explore the function of sgRNA: In the context of genome editing, sgRNA (single-guide RNA) is used to guide the Cas proteins to the specific DNA sequence that needs to be cut. However, in bacterial cells, the natural function of CRISPR is to protect against viral DNA, not to detect DNA to cut.
Examine homologous recombination: Homologous recombination is a natural process used by cells to repair broken DNA. It is not always used in genome editing, as other methods like non-homologous end joining (NHEJ) can also be employed.
Consider the possibilities of genome editing: Genome editing can be used to modify genes, such as inserting new sequences, as well as to disrupt them, which involves creating mutations or deletions.
Review each statement based on the explanations: Use the information from the previous steps to determine the validity of each statement regarding genome editing.
 Verified video answer for a similar problem:
Verified video answer for a similar problem:This video solution was recommended by our tutors as helpful for the problem above
Video duration:
2mPlay a video:
Was this helpful?
Key Concepts
Here are the essential concepts you must grasp in order to answer the question correctly.
Cas Proteins as Endonucleases
Cas proteins, particularly Cas9, function as endonucleases in genome editing. They are enzymes that can cut DNA at specific sites, guided by RNA molecules. This cutting ability is crucial for the CRISPR-Cas9 system, allowing precise modifications or disruptions of target genes.
Recommended video:
Guided course
Proteins
sgRNA in Genome Editing
Single-guide RNA (sgRNA) is a synthetic RNA molecule used in genome editing to direct Cas proteins to specific DNA sequences. In the CRISPR-Cas9 system, sgRNA binds to the target DNA and guides the Cas9 enzyme to the correct location for cutting, enabling precise genetic modifications.
Recommended video:
Guided course

Genomes and Genome Evolution
Homologous Recombination in DNA Repair
Homologous recombination is a natural DNA repair mechanism that uses a homologous sequence as a template to accurately repair breaks. While it is a common method for repairing DNA, it is not always used; cells may employ other repair mechanisms like non-homologous end joining, depending on the context and available resources.
Recommended video:
Guided course

DNA Repair

 5:30m
5:30mWatch next
Master Genomes and Genome Evolution with a bite sized video explanation from Jason
Start learningRelated Videos
Related Practice